You are here:
MRST
/
Modules
/
MRST-co2lab
/
Vertical-Equilibrium Models
/
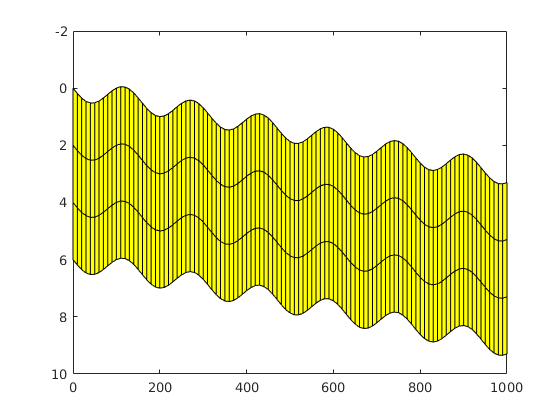
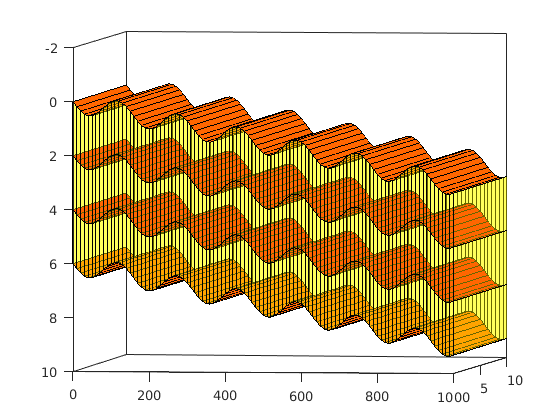
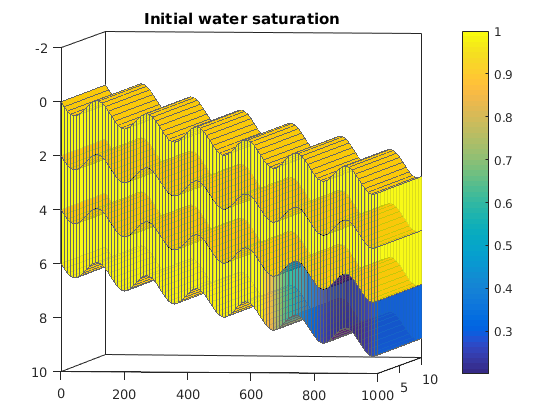
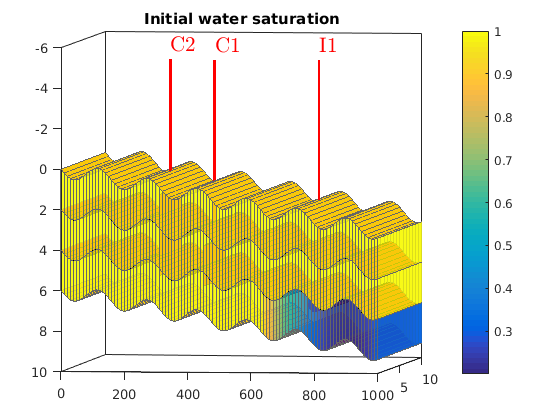
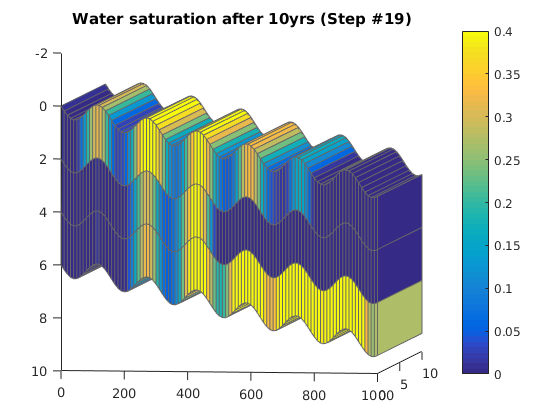
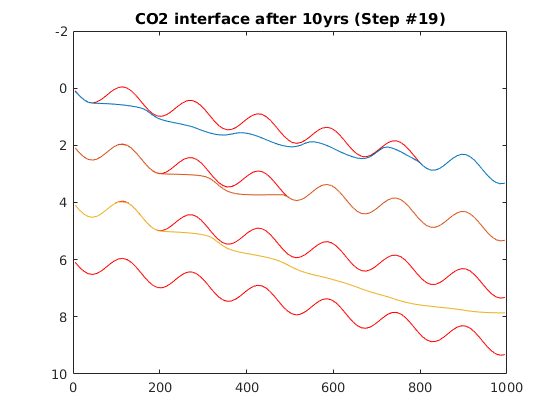
Simulating Leakage in a Multilayered System